| |
Annie organizes experiments like we were taught in school - keep a notebook with numbered pages, date everything, title all your pages with the experiment name, etc. And of course in the computer world, back everything up! Twice! Annie does many of the mundane chores that clutter up scientific and engineering workflows, and she'll adapt to your style and your preferences.
Let's look at how Annie works.
When defining a simulation, you have to give it
a name. The syntax is: SIMULATION < simulation_name >
Next you have to tell the simulation which network(s) to use, you use one
or more lines like the following: USING NETWORK RETINA
Templates are files that define stimuli. Stimuli are geometric patterns with
relative coordinates (relative to the base of the template). For example in the
visual field, there are points of light, bars, circles, annuli, gratings, and
so on. These stimuli can move, which is what the "apply" defines, it defines
how the template moves. The stimulus template is "applied" to a particular
cell group (a cell group is part of a network, defined below). The syntax is
very simple: APPLY TEMPLATE < template_name > TO < cell_name >
You can load the template files up front, with one or more directives of
the form LOAD_TEMPLATE_FILE < filename >
Within each template file are one or more template definitions. These are
named. In the APPLY statement, you use the template name. (Not the file name).
Each APPLY statement has multiple directives that control the action of the
template on the designated cell group. Template actions are additive, for
example if you have two dim spots of light affecting the same cell, they
will add together to form a larger response. (Template actions can be made
inhibitory using the SCALE_LEVEL directive which is discussed below
in the section on templates).
Here is an example of a complete simulation definition file. Any line that begins with an asterisk is a comment.
SIMULATION RETINA_SIMULATION
* if there's no USING statement there can only be one network
USING NETWORK RETINA
* template names are defined in the template file
LOAD_TEMPLATE_FILE cones.tem
* apply a named template to a cell group
APPLY TEMPLATE T1 TO LIGHT
STARTING_COORDS (-9000,-9000,1000)
MOVEMENT_COORDS (1000,1000,0)
SCALE (2,2,0)
START_TIME 0
STOP_TIME 12
INCR 2
LOOP 7
REPEAT FOREVER
The coordinates are defined relative to the network. The network coordinates
form a "coordinate space" within which neurons are populated and within which
templates are applied. The coordinate space is defined by the network.
You don't have to have a USING NETWORK directive, if there's only one network
it doesn't have to be explicitly declared. However simulations can use more than
one network. (Like, dorsal and ventral visual streams). A network is a convenient
conceptual abstraction, is has no real significance other than being named and
defining the coordinate space. It serves as a "container" for other objects,
like neurons and synapses. Networks are discussed below, let's return to the
stimulus and template definition for a moment. Simulation definition files
have a .SIM extension.
Each template has a size, which defines its extent relative to (0,0,0).
Stimuli are simple elements, like POINT, BAR, CIRCLE, ANNULUS and so on.
These simple elements are put together to form more complex stimuli, for
example gratings are combinations of bars. POINTS just have locations
and levels, but BARS have beginning and ending points that define orientations.
Circles have diameters, they're essentially like points with extent. Annuli
have inner and outer diameters. Annie knows about triangles, rectangles, polygons of all kinds, and significantly, meshes. Annie has an enormous geometric library for three dimensional generation and manipulation of everything from layered and modular nuclei to fiber tracts to the synapses themselves.
To use a template, you declare it inside a template file. Each template
has a name and a size, as follows: TEMPLATE < name > SIZE (x,y,z) where
x, y, and z are template coordinates (not necessarily related to any
network coordinates, they are mapped on the basis of extent and scale).
Any coordinate can be 0, for example if you have a 2-dimensional visual
grating you can define it as SIZE (1000,1000,0), or SIZE (1000,0,1000)
if you want it applied in the x-z plane. Other orientations are possible, using the ORIENTATION keyword.
The above example is a moving template, it has MOVEMENT_COORDS (1000,1000,0)
and an INCR 2 which means it will execute the movement every 2 msec.
If you leave out the INCR statement the movement is executed every
msec. Annie's default tick rate is 10 ticks/msec, so if you specify
an INCR of 1 (or just omit the declaration), the template will move
every 10 ticks on the screen.
Template elements ordinarily just have levels, but they can also have
functions attached to them (which determine the levels). If you're using
a simple level, you can just say LEVEL < level >, and you also have the
option of using a SCALE < scale > (on the same line) or a LEVEL_SCALE < scale >
(on a separate line) which multiplies the level. This is useful when
you need an inhibitory template, for example a visual stimulus representing
"dark" relative to the background.
Template definition files have a .TEM extension. Here is a complete
example of a template definition file:
TEMPLATE T1 SIZE (6000,6000,0)
BAR (0,0,0) TO (2000,1000,0) LEVEL 5.0
BAR (3000,3000,0) TO (5000,5000,0) LEVEL 5.0
TEMPLATE T2
* SIN(A,K,W,P) in radians
POINT (9250,9000,0) FUNCTION SIN(10.0,0,10.0,0)
TEMPLATE T3
CIRCLE (4000,4000,0) DIAMETER 2000.0 LEVEL 5.0
TEMPLATE T4
ANNULUS (3000,3000,0) OD 2000.0 ID 1500.0 LEVEL 5.0 LEVEL_SCALE -1.0
The first example shows a pair of simple bars (one of which is a square).
The second example shows the use of a function to determine the level
(note there is no level directive here, the function sets the level).
Functions are internal functions, they must be known to the system.
The list of available functions is below. The last example shows a
LEVEL_SCALE directive, making the annulus dark relative to the
background. When this stimulus is applied, a value of -5.0 will be
transferred to the target cell.
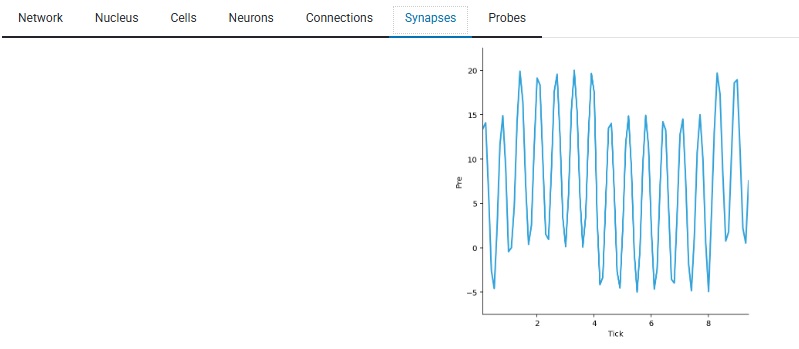
Here is the effect of template T2 (above) on a synapse. You can see the sine wave in action, and in this experiment a nearby slightly weaker grating of light is also affecting the synapse, you can see the modulation as it moves across the cell's receptive field.
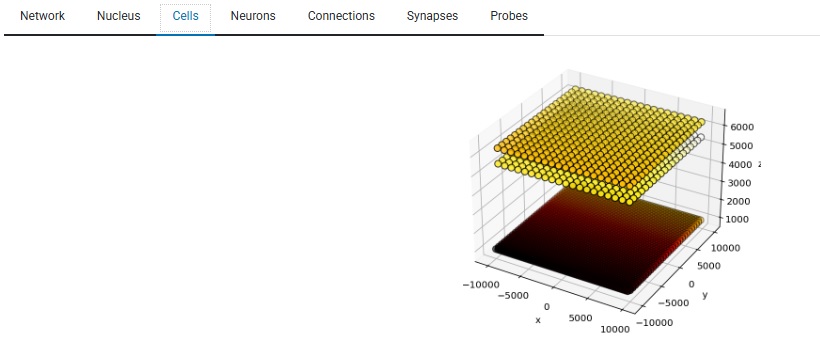
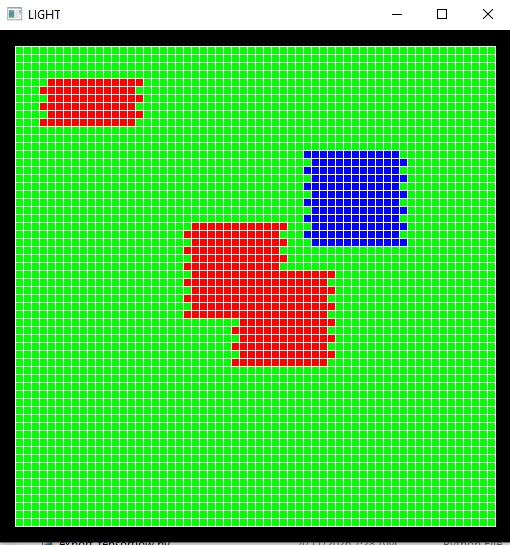
By the way, the graphics are highly interactive in a live Annie environment. You can zoom in and out, rotate the view, save any view to a file, that kind of thing. If you're in the simulation business, visualization is a big deal. For instance, a time series of synaptic activity doesn't really help me understand receptive fields, what I really need to see is the pattern of activity in a cell group, relative to the stimulus input. Here is one of many ways Annie can visualize this for you. You can look at your cells as a grid, even if they're not geometrically arranged that way, and if you have a topographic mapping you can see how the stimulus maps onto the network activity in real time.

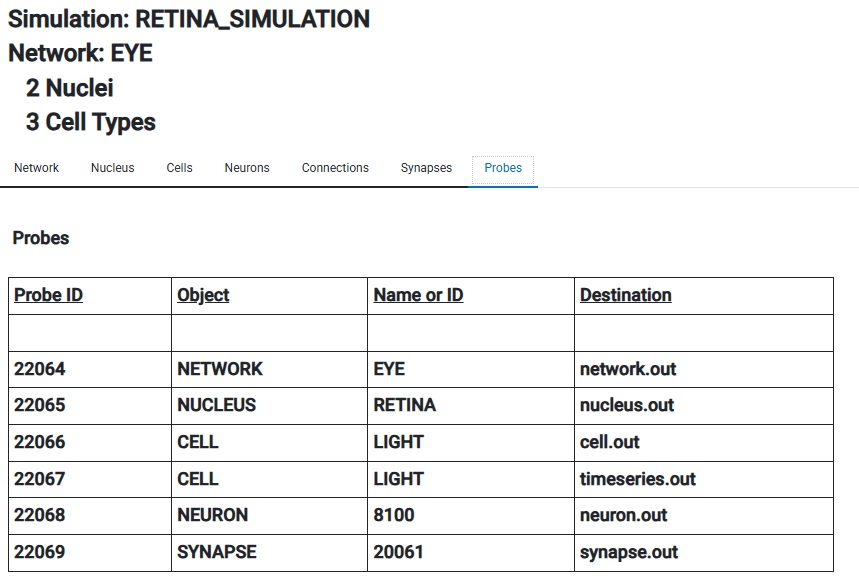
And of course you can still see information the old fashioned way too. Here are some "probes", which are ways of getting information out of Annie. In a personal context, data files are the most common need, but in the full multiprocessing client-server version of Annie you might see entries on the right that say "Client 17 Window 5", and those would be real-time streams destined for a workstation.
Now, on to the subject of networks. Annie uses the same simple philosophy to define networks and be able to organize them and re-use them in sequences of experiments. No programming is required, only the ability to tell Annie what you need. There are some very sophisticated visualization tools available in the data science community these days, and Annie can export a CSV file containing a time series for every neuron in a cell group, that can be read directly into Pandas without modification and visualized using matplotlib, Plotly, HoloViews, or any of your favorite toolsets. You can do that with one line in your .SIM file:
PROBE CELL blue_cones TYPE level FILE timeseries.out
This will apply a probe of type "level" to the entire cell group called "blue_cones", and save the results in a file called "timeseries.out". The syntax is PROBE < probe_what > TYPE < probe_how > < output_type > < send_output_where >. There are additional parameters with which you can control probe durations (they're just like templates, you can apply them to objects, starting and ending at certain times). You can probe activity, geometry, and many other things, and Annie will create high level "views" if you wish, to save on data transfers and speed up simulations.
Besides the stimuli which were defined above (organized into a useful
form on the basis of libraries of stimulus templates,
which can be created and edited using Annie's template editor, or Notepad if you prefer), there are
two parts of a simulation, the geometry, and the engine itself. Annie
addresses a gap between large highly structured algebraic simulators like
TensorFlow, and single-cell simulators like Brian2 that require you to
write differential equations. Annie thinks like a neuroscientist, not
like a machine learning engineer. (If you want machine learning, use
TensorFlow). Annie has amazing geometric capabilities. Annie allows you to define your network directly in terms
of brain anatomy and physiology. Annie knows about nuclei, neuropils,
modules, layers, and synaptic capsules. Annie has a sophisticated
geometry engine that creates axons and dendrites and properly inserts
them into their geometries and connects them with their destinations. You don't have to provide Annie with connection maps, she already has them. If you have a topographic connection from one layer to the next, you can say DIVERGENCE 5 and you're done, Annie will connect each cell to its 5 nearest neighbors in the destination layer. No fuss, no muss. Easy.
To illustrate a NETWORK definition, we can use the human retina. It's a great
example because all brain areas are this way, you can model them as
simple or as complex as you want.
In a simple model, a human retina could have 3 layers of feed-forward
connectivity (photoreceptors, bipolar cells, and ganglion cells), and
2 lateral paths (horizontal cells and amacrine cells). In a slightly
more sophisticated model the lateral paths could be organized into
neuropils (inner and outer plexiform layers), and Annie can be told
about the shapes of axons and the branching patterns of dendrites
(and can generate them consistently, or pseudo-randomly with variation
according to specified distributions). In a full-blown model the
5 layers become 10, and the plexiform layers are further sub-divided,
and there are 12 types of bipolar cells and 60-ish types of amacrine
cells and over 20 different kinds of ganglion cells. Furthermore, the
horizontal cells form an electrical syncytium via gap junctions, and
this behavior is essential for the creation of the "surrounds" of
bipolar cell receptive fields, which are usually more than 20 times
larger than the corresponding extent of a horizontal cell dendritic
tree. So even this simple structure gets complex real fast, if we
wish to model it that way. Maybe we're only interested in the
dynamic response of ganglion cells, but maybe we're interested
in the contribution of Muller cells to the retinal B-wave, and
maybe we want to study the effect of electrical waves generated
by amacrine cells during development.
To support the rich anatomy of the human brain, Annie has a sophisticated
anatomical model, and a large library of available geometries to handle
the majority of the structures encountered in current research. At a high
level this includes things like nuclei and sub-nuclei, projections and
fiber bundles, layers and modules with specific connectivity, synapses
with a wide variety of arrangements (including capsular), axonal and
dendritic compartmentation, and so on. To maintain compatibility with
other simulators, Annie understands the industry-standard neural models
and the differential equations and numerical methods underlying them.
In most cases these can be invoked with very simple declarations in
the network definition file.
Annie likes Occam's Razor. Simpler is better, because simulations are complicated enough. But there are some essential ingredients to being able to run experiments. Many scientists spend a small fortune on software to orchestrate stimulus presentation. Whereas Annie is open source and free for personal use. Annie is written in Python, has only a few graphical dependencies, and will run just about anywhere (not on Termux in a cell phone, but anywhere else). You can use Annie like a Python library if you wish, the API is published in detail. Or, you can just say "pip install annie-interface" and "annie" to start Annie's Personal Neural Network Simulator.
Far Out - Show Me How To Build A Network
| |



